“…Although it is true that the PhD degree has been criticized in recent years for remaining classical and reluctant to being seduced by a more market-oriented skill-based training system, it still is the strongest higher-education formative experience ever developed in academia. The globalization of science and technology and international exchanges of professionals have led to agreements that attempt to equate higher-university titles across disciplines (the functional equivalents to a PhD), but it has also triggered worldwide the proliferation of fake, self-granted PhD acronyms to doctoral degrees that are no match to the education provided by the rigorous PhD-granting institutions…”
by Guillermo Paz-y-Miño-C
Before listing what makes a PhD-training unique, I will share the latest data about the United States. The statistics below come from the National Science Foundation and correspond to 2016 (released early December 2017). You will have to wait until late 2018 for NSF to process the data from 2017.
 In 2016, the United States graduated 54,904 PhDs and doctorates (the latter included non-PhDs but doctors in education, law, business administration, social work, international relations, health-care professions like physicians, veterinarians, dentists, nurses and clinical psychologists, and technical posts like pharmacy) at 436 higher-education institutions. Of these graduates, 38,406 were American citizens and residents, and 16,498 were temporary visa holders of diverse nationalities (=international students). The top five PhD/doctorate-granting institutions were the University of Texas-Austin (849), University of Wisconsin-Madison (823), University of Michigan-Ann Arbor (819), University of California-Berkeley (796) and University of Minnesota-Twin Cities (787).
In 2016, the United States graduated 54,904 PhDs and doctorates (the latter included non-PhDs but doctors in education, law, business administration, social work, international relations, health-care professions like physicians, veterinarians, dentists, nurses and clinical psychologists, and technical posts like pharmacy) at 436 higher-education institutions. Of these graduates, 38,406 were American citizens and residents, and 16,498 were temporary visa holders of diverse nationalities (=international students). The top five PhD/doctorate-granting institutions were the University of Texas-Austin (849), University of Wisconsin-Madison (823), University of Michigan-Ann Arbor (819), University of California-Berkeley (796) and University of Minnesota-Twin Cities (787).
The top five PhD/doctorate-granting institutions to the international students were Purdue University-West Lafayette (372), Texas A&M University-College Station and Health Science Center (328), University of Florida (297), University of Illinois-Urbana Champaign (278) and Ohio State University-Columbus (267).
23% of the graduates obtained degrees in the life sciences, 17% in engineering, 16% in psychology and social sciences, 12% in physical and earth sciences, 10% in humanities and arts, 9% in education, and 7% in mathematics and computer sciences; the rest graduated in other fields, including business, management and administration, as well as communication (adding to 6%).
54% of the graduates were men and 46% were women. The men/women gap varied across fields, as follows: in the life sciences, 55% were women and 45% were men; in engineering, 77% were men and 23% were women; in psychology and social sciences, 59% were women and 41% were men; in physical and earth sciences, 69% were men and 31% were women; in humanities and arts, 52% were women and 48% were men; in education, 70% were women and 30% were men; and in mathematics and computer sciences, 76% were men and 24% were women.
“All PhDs are doctoral degrees, but not all doctoral degrees are PhDs”
What did those who got PhDs do to be granted the degrees of “philosophiae doctoris” or “doctor of philosophy”? My listing below is summarized and applicable particularly to the life sciences:

“…if your doctoral certificate does not read ‘philosophiae doctoris’ or ‘doctor of philosophy,’ chances are you might not have a PhD…”
Number 1 – Application. All began when they sent applications to multiple PhD programs in the US, knowing that, if lucky, a couple of institutions might accept them. Crucial to their applications were five components: a statement of interest (a sort of letter of intent, but broader, explaining why they wanted to pursue research), the curriculum vitae (CV), three letters of recommendation (preferably by academics with PhDs themselves), GRE / TOEFL scores if already available (the originals were sent directly to the universities by the GRE– / TOEFL-test agencies), and the undergraduate academic records (the “transcripts”). The personal statement had to be competitive in relation to the hundreds/thousands of statements submitted by students from all over the world. It revealed, in a few pages, the genuine intellectual potential of the applicant; his/her curiosity-driven mind and interest in seeking scientific knowledge. The CV needed to summarize the evidence that the applicant was a scientist in the making (format and content had to be just right). The letters of recommendation convinced a reviewing committee (made of professors in multiple fields and student representatives to the “graduate committee”) that the applicant will endure the challenges of the PhD academic environment (survive and succeed in it). The undergraduate transcript had to simply document that the applicant was as ordinarily outstanding as the other applicants (previous research experiences were always a plus, even more important than the excellent grades). Perhaps now the reader realizes how significant were the personal statements and CVs (the latter as supplements), considering that the letters of recommendation and undergraduate records of all competitive applicants were comparable.
Number 2 – Interview. This was done in multiple ways and more than one semi-formal or formal interview(s) took place. Via Skype (the first approach) and a later visit to the graduate program during which the potential student met with the graduate committee (usually with each of its members and also with the group), interacted with diverse research teams with which the student might work in the future (if accepted into the program), participated in a journal club discussion or laboratory meeting (for which the applicant was given scientific publications to read in advance, so that his/her contribution to the discussion was meaningful), and socialized over dinner, or equivalent gathering, where members of the program (faculty, postdocs, graduate and undergraduate students) got to talk, briefly, to the applicant. As the interview(s) made progress, the members of the graduate committee got feedback from those whom interacted with the applicant(s). Each potential student was ranked in respect to others and, after various meetings, the graduate committee made recommendations (to higher-instances in the institution) as to the list of students whom should be offered a “graduate student line” (i.e. the type of funding the student shall receive, either as research or teaching assistant, or both). In most cases, a database was created with quantitative scores as per all aspects relevant to the applicant(s) performance: personal statement, CV, letters of recommendation, undergraduate transcripts (this also applied to those with masters degrees in a separate column), GRE/TOEFL scores (or equivalents), research experience, publications (when applicable, as coauthor or leading author), presentations at regional, national or international scientific meetings (posters, talks), scientific competitive awards (including mini-grants), interview performance, participation in journal club/laboratory meeting discussion while visiting the program, and interaction with members of the department (e.g. scoring 1 = poor; 2 = fair; 3 = good; 4 = very good; and 5 = excellent). All applicants were ranked from highest to lowest scores, and only those voted positively by the graduate committee were contacted and offered the graduate student line. Not all the top-ranked accepted since some had several offers.
“…during two years, the PhD students were transformed by a rigorous academic environment in which ‘quality education’ was at the center of their intellectual development…”
Number 3 – The First Two Years. The entire PhD program lasted about 6 years; but years 1 and 2 probably marked the most significant transformation in the students. They were required to enroll in courses that emphasized further development of their analytical skills (i.e. strong in statistics, data processing or synthesis-writing), integration of information from the primary literature (hundreds of articles were required to be read) and weekly take-home assignments (mini-reviews of the literature and/or long papers addressing conceptual questions prepared by a professor). If the students had previous masters degrees or non-PhD-doctorates, only a fraction of the courses already taken were accepted into the PhD program. Everyone took new courses taught by world specialists in specific fields; double dipping was rarely allowed in more than 1/4 or 1/3 (exceptionally 1/2) of the total PhD curriculum. — Thus, in most cases, two in-house courses were taken per semester, combined with a discussion-based-third-course (a “graduate seminar”) based on weekly student presentations of scientific articles (about 5 articles per student, per week, in a class of 16 students and 1-2 instructors/facilitators). The latter were the foundation of the training in the “argumentative format,” which the students got to internalize. The graduate seminars were both in-depth academic discussions of the philosophical foundations (as per philosophy of science) of recent or classical literature, and intense exchanges in which the student leading the session was grilled by his/her peers and instructor(s). This format helped students to become aware of their academic strengths and weaknesses; all in the open, with peers and mentors watching. — Many PhD students worked 14 to 16 hours a day, all days; some took on and off naps while working continuously for weeks, occasionally months under that rhythm. At public institutions, most PhD students had “teaching-assistantship” lines (TAs) that paid their salaries, and they were responsible for teaching or co-teaching laboratory or reading sessions for freshman/sophomore undergraduates, grading exams and reports, and holding weekly office hours to mentor undergraduates. Each semester, their teaching performances were evaluated by the undergraduates and TAs-supervisor(s). — On top of the course work and TA responsibilities, all graduate students were required to attend weekly seminars by an internationally-known speaker invited by the program; they met with the speaker one-on-one, or in small groups, and read articles published by him/her. Weekly, sometimes twice a month, members of the PhD program (faculty or graduate students) hosted at their homes/apartments semi-formal evening-talks by local speakers (from sister institutions in the area); these discussions were enthusiastically attended by the “graduate community” (a voluntary practice built on the desire to reinforce a culture of learning). — And on top of these activities, the students participated at weekly meetings with their PhD advisor’s team in which they discussed the science being done in the laboratory or research group. At such meetings, the fresh PhD students were constantly taught by everyone else in the team (i.e. advanced research undergraduates, other PhD students, postdocs and the principal investigator) the methodological and conceptual rationale behind the research carried out by the team. — In many programs, the students participated in mandatory “rotations,” which consisted in doing practical research work at a laboratory, or with a research team other than his/her own and, after a semester, presented the results of that experience to the entire academic department in a public talk (1 to 3 rotations during the first two years of the PhD program, each up to 20 hours/week work). The purpose of each rotation was to expose the students to different fields of scientific inquiry, diverse working environments and mentors, and open the possibility for the students to change their minds and complete their PhDs in fields just discovered during the rotation experiences. — Some programs also required the students to participate in practical internships (3 to 8 weeks), during the summer of their first or second academic years (paid by the PhD program or by an external agency). As interns at known State or Federal agencies, or at NGOs, the graduate students “tasted and practiced, hands-on, the professional world” (e.g. National Institutes of Health, US-Congress, The World Bank, Centers for Disease Control and Prevention, National Geographic, Department of Justice, Smithsonian Institution, Department of Labor; click on internship opportunities for graduate students). At their return, they submitted reports of their experiences and presented talks to their departments. — In sum, during two years, the PhD students were transformed by this rigorous academic environment in which “quality education” was at the center of their intellectual development.
“…Only if the students passed the qualifying exams satisfactorily, they were finally and officially called ‘PhD candidates,’ a placebo status of psychological value only to students…”
Number 4 – Qualifying Exams (Preliminary Exams = Prelims, or Candidate/Candidacy Exams). Before the second year was completed, PhD students took the most challenging test ever in their lives. The goals of the qualifying exams were multiple: test the analytical-thinking ability developed since the students joined the PhD program; assess the academic maturity gained during the first two years; evaluate their retention of information and capacity to integrate scientific knowledge coming from multiple sources; verify their capacity to answer “why” questions (i.e. ultimate causality) when confronted with theoretical academic scenarios that they were asked to solve; challenge them to propose the immediate and future directions an entire academic field should take to make significant transformations in science. These goals were accomplished via different exam-formats, including complex questions given to the students in advance, which they had an entire semester to think about, compile library information, scrutinize scientific papers and structure comprehensive answers to later be presented in written and/or oral examinations. Other formats consisted in developing research proposals on topics unknown to the students, although tangentially related to their areas of expertise (according to NIH, NSF, Department of Education guidelines), and that the students had to prepare as if they were world experts and later defend such proposals in front of the graduate committee. Or prepare a theoretical review of the literature, in an entire field of expertise, and present it to and answer questions from a team of professors. The qualifying exams took months to prepare (including in most cases the end-of-the-year break –no vacations!) and 1-2 entire days of in-writing and/or oral examinations by the graduate committee. — Only if the students passed these exams satisfactorily, they were finally and officially called “PhD candidates” (PhDc), a placebo status of psychological value only to students. In many programs, those who did not pass the qualifying exams were given a terminal masters degree and sent home. A few programs granted the students a second chance to retake the exams (usually in a more rigorous setting) and made final decisions by the end of the second academic year. — Some programs did not require qualifying exams, but had other formats of PhD-candidacy assessment; such programs were exceptions, rather than the norm.
Number 5 – PhD Dissertation Proposal. This document had to be approved by the students’ PhD-dissertation committees, and later defended in a public presentation (sometimes in close-door meetings with the thesis committees) at the end of the first or second semesters of year 3. The “thesis proposal” was comprehensive in its theoretical background (placing the students’ intended research in a historical context, as per the chronology of a research field), with central and auxiliary hypotheses to be tested, detailed in methodology, statistical analyses, expected results (i.e. figures and tables already built and specifying all outcomes based on hypothetical data), significance of the work in terms of generating new knowledge and advancing science, future grant-proposals to be submitted to local, regional or national agencies, as well as the forecast of conceptual publications (at least three) to be generated from the research. — The thesis committee (per individual student) likely included five members: an advisor, sometimes a co-advisor, 2-3 professors from within the PhD program, and a researcher from another PhD-granting institution. Depending on the field of expertise and program, the thesis committees occasionally included even more members (as many as the students needed for proper advice; 6-8 members were not rare). The approval of the dissertation proposal was done first by the thesis committee and later certified by the Department (remember that faculty, postdocs, graduate students and research undergraduates attended the oral presentation and thesis-proposal defense) and university. The officially approved thesis proposal turned into a “contract” that the students had to complete satisfactorily, within 3-4 years, in “partial fulfillment of the requirements for the degree of” philosophiae doctoris or doctor of philosophy. Note that some programs granted the PhDc status only after the thesis-proposal was approved (i.e. after the first two years of course work plus completion of the qualifying exams); thus, the students became PhDc during year 3.
“… the science was conducted in the open, shared with the institutional community, and subject to criticism and feedback all the time… Advanced or just starting in their PhD education, the students were pushed beyond their comfort zones. Seniority was rarely observed…”
Number 6 – The Research, Grant-Proposal(s) Submission(s), Participation at National and International Meetings. During years 3, 4 and 5 the students conducted the research to which they committed themselves in the PhD-thesis proposal. Those in TA lines continued to teach. — Weekly or monthly, they reported progress to their advisors and research teams (semi-formal oral presentations of results and/or difficulties delaying the project); once per semester, they met with the thesis committee for the same purpose (i.e. semi-formal presentation of data and analysis of partial results); and once a year they submitted comprehensive progress reports to the PhD Program Director (i.e. the parts of the research already completed, work in progress, poster or oral presentations at significant scientific meetings, awards, mini-grants or grants under review, as well as those that were funded) and received a letter of evaluation from the PhD Program Director; in it, the individualized expectations of the program to secure timely completion of the work were specified (note that the continuation of institutional funding to the student’s salary, including the TA line, depended on a positive evaluation). — As PhD-researchers, the students were expected to train undergraduates and provide and receive feedback to/from peers (this was part of the culture of academic reciprocity, which the program community embraced). They were also required to submit doctoral-dissertation grants to national agencies (e.g. NSF), regardless of them being funded or not. Each semester, they submitted competitive mini-grants to their own PhD-programs (from less than $1000 to up to a few thousand dollars), departments, colleges or universities, as well as to scientific societies (i.e. funds for traveling to international meetings or for materials and logistics related to the research). Each of those grants varied in narrative-length and conceptual/practical emphasis; in this way, the students became skillful at summarizing their projects in a few hundred words, or in lengthy documents with detailed budgets and justified expenses; all on a competitive basis and repetitively during years 3-5. Those researching off campus, in the field or other countries, returned each semester, or yearly, to their institutions and shared their progress via oral presentations (data oriented) and discussions with the entire program. Thus, the science was conducted in the open, shared with the institutional community, and subject to criticism and feedback all the time. — While on campus, the students continued to participate in journal clubs, seminar discussions, advising and mentoring sessions. Advanced or just starting in their PhD education, they were all pushed beyond their comfort zones. Seniority was rarely observed.
“… Every PhD dissertation aimed at becoming a unique and crucial contribution to science (work that had never been done before); many, perhaps most, accomplished it…”
Number 7 – Completion of Data Collection, Analysis and Writing of the Dissertation. By year 5 in the PhD program, the data collection ended and both the comprehensive statistical analyses and writing of the dissertation began. This was likely done in combination with a couple of papers already published, or manuscripts submitted during years 4 and 5 (or extended to year 6), but many students could not write such papers until full collection and processing of the data. It did take from six months to a year to complete the no-long-ago seemingly-eternal thesis. In the natural sciences, engineering and mathematics, the thesis was fairly short (200-300 pages, including raw data and analyses attachments), but in the humanities and arts the documents often reached +400 pages. Depending on the field of specialization, the students were expected to publish, at least, three comprehensive, conceptual (not descriptive!) papers of their own that contributed significantly to advance the scientific knowledge in their fields. As members of a productive team of researchers, they likely co-authored an equal number of additional articles (a combination of descriptive work, notes, reviews with their advisors, or joined papers with master students or undergrads). The important aspect of the PhD research was that it covered interrelated, yet separate topics (usually three, but often four or five), each comprehensively designed for testing, with specific conceptual questions, hypotheses, predictions and exhaustive approaches to examining each hypothesis and its predictions. Every PhD dissertation aimed at becoming a unique and crucial contribution to science (work that had never been done before); many, perhaps most, accomplished it.
“…After a month or so, the graduates received in the mail a modest certificate declaring them ‘philosophiae doctoris’ or ‘doctor of philosophy.’ Another document of little use considering that the certification that a PhD degree had been granted was usually extended by the university via official copies of the graduates’ transcripts –with the entire academic history…”
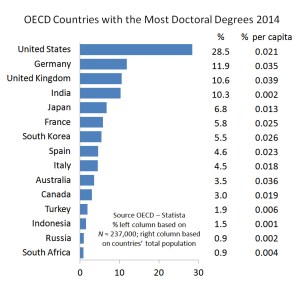
Percentages of doctoral degrees granted by OECD countries in 2014; left = N ca. 237,000; right = per capita (click to enlarge).
Number 8 – Thesis Defense and Graduation. The departmental seminars in which the PhD candidates presented and defended their dissertations were announced publically and attended by faculty from various universities, postdocs, graduate/undergraduate students and, occasionally, by the candidates’ family members. The students were introduced as “today’s seminar speaker” or, more frequently, in non-solemn manners since, although successful presentations and thesis-defense were expected (otherwise the students had not been allowed to get that far in their programs), prudent enthusiasm was honored. The 50-minute seminar was followed by 30-40-minutes of Q&A by the audience, and by close-door meetings between the students and their graduate committees (up to 2-3-hours). By the end of the day, social gatherings were often organized to celebrate the events. During the following weeks, sometimes months, the students made adjustments to the dissertations (from minor to substantial) and prepared final versions of the documents. These needed signatures of approval by each committee member. The properly formatted original and copies of the documents were then submitted for the universities’ final approval and filing. [Note that copies of all PhD dissertations written in the US are stored at national archives with access to the public, examples include ProQuest or DissExpress, but there are others]. Attendance to “graduation day” (i.e. the universities’ bi-annual ceremonies) was an option for all PhD students; quite a few did not participate (a common excuse was their mental fatigue and desire to just move on). After a month or so, they received in the mail a modest certificate declaring them “philosophiae doctoris” or “doctor of philosophy” (another document of little use considering that the certification that a PhD degree had been granted was usually extended by the university via official copies of the graduates’ transcripts –with the entire academic history).
To close: the National Science Foundation indicates that, in 2016, about 30% of the PhD/doctorate graduates in the life, physical and earth sciences were headed to postdoc positions (2-3-years of additional research experiences in science productivity). By contrast, only 17% of their counterparts in mathematics, computer sciences and engineering had similar plans. And about 60% of most graduates were simply moving into research and development jobs (see Science).
Although it is true that the PhD degree has been criticized in recent years (see A, B, C, D) for remaining classical and reluctant to being seduced by a more market-oriented skill-based training system, it still is the strongest higher-education formative experience ever developed in academia. The globalization of science and technology and international exchanges of professionals have led to agreements that attempt to equate higher-university titles across disciplines (the functional equivalents to a PhD), but it has also triggered worldwide the proliferation of fake (see E, F), self-granted PhD acronyms to doctoral degrees that are no match to the education provided by the rigorous PhD-granting institutions. — EvoLiteracy © 2018
You can contact Guillermo Paz-y-Miño-C via email at guillermo.pazyminoc@gmail.com — Follow us on Twitter @gpazymino and Facebook.
Related Articles
Intolerance to Free Speech at America’s College Campuses
College Educated But Deeply In Debt For An Overpriced Degree
Imminent Collapse of Basic Science Under For-profit Model
Dehumanizing Academia by Dismantling the Humanities
Fragmentary Truths and the Intellectual Imbalance in Academia
























You must be logged in to post a comment.